Professor Anna Philpott
Websites
- CRUK Cambridge Centre: http://www.cambridgecancercentre.org.uk/users/annaphilpott
- Cambridge Neuroscience: http://www.neuroscience.cam.ac.uk/directory/profile.php?ap113
- Wellcome Trust Medical Research Council Stem Cell Institute: http://www.stemcells.cam.ac.uk/researchers/principal-investigators/anna-philpott
Graduate Students
Professor Anna Philpott is not currently taking on graduate students.
Research Interests
Cell Fate and Cell Cycle in Development and Disease
Cell fate and cell differentiation changes across developmental time and is usually co-ordinated with changes in cell proliferation. However, mechanisms that allow cells at progressive stages of development to respond differently to the same fate determinants and mechanisms that link the cell cycle and differentiation are poorly understood. Our laboratory is interested in understanding how cell fate choices are made and how cell cycle is co-ordinated with cell fate determination and differentiation using early Xenopus frog embryos, as well as in ES cells, organoids and mouse models.
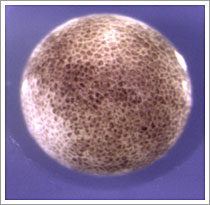
This image on the right is a blastula stage Xenopus embryo showing multiple, synchronously dividing cells and these cells are making sophisticated choices largely based on the presence and activity of transcriptional activators whose function depends on the state of the chromatin of the cells in which they are found. We aim to understand how transcriptional regulators of cell fate and differentiation interact with the chromatin and cell cycle environment to confer cell behaviour. We focus particularly on development of the embryonic nervous system, of mesoderm and of endocrine lineages in the gut and pancreas, where fate decisions and the decision to divide or differentiate is controlled by the activity of proneural transcription factors.
Recently, we have been investigating links between the core cell cycle machinery and mechanisms of differentiation. We see that cell cycle components directly control several modes of post-translational regulation of these proneural transcription factors, which in turn, regulates the balance between precursor cell maintenance and differentiation. We are now using genome-wide approaches in multiple and single cells to further determine what controls the ability of cells to respond to lineage specifiers in development and reprogramming.
Moreover, we are also interested in dysregulation of these mechanisms in cancer cells. Our work in embryonic development is revealing how cells stop dividing and adopt a normal differentiated fate, and is pointing to ways that cancer cells lose their differentiated phenotype and re-enter the cell cycle precociously during tumourigenesis.
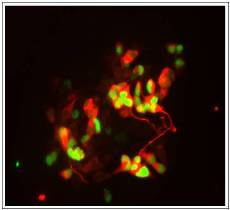
In particular, we are currently attempting to manipulate these processes to promote cancer cell differentiation to effect remission, particularly in neurologically-derived tumours such as neuroblastoma and glioblastoma.
The image on the right shows embryonal carcinoma cancer cells can be induced to differentiate into neurons in vitro by expression of a proneural transcription factor that has been mutated so it can no longer be phosphorylated by cyclin-dependent kinases. GFP marks cells expressing the proneural protein, while a marker of neuronal differentiation is detected in red.
Publications
Pubmed journal articles - pubmed
Key publications
Neurogenin3 phosphorylation controls reprogramming efficiency of pancreatic ductal organoids into endocrine cells. Azzarelli R, Rulands S, Nestorowa S, Davies J, Campinoti S, Gillotin S, Bonfanti P, Göttgens B, Huch M, Simons B and Philpott A. (2018) Scientific Reports. Oct 18; 8(1):15374. doi: 10.1038/s41598-018-33838-5.
Phospho-regulation of ATOH1 Is Required for Plasticity of Secretory Progenitors and Tissue Regeneration. Tomic G, Morrissey E, Kozar S, Ben-Moshe S, Hoyle A, Azzarelli R, Kemp R, Chilamakuri CSR, Itzkovitz S, *Philpott A and *Winton DJ. (2018). Cell Stem Cell. Sep 6;23(3):436-443. doi: 10.1016/j.stem.2018.07.002. * Joint corresponding authors.
The developmental origin of brain tumours: a cellular and molecular framework. Azzarelli R, Simons B and Philpott A. (2018). Development, 145,145(10). pii: dev162693. doi: 10.1242/dev.162693
Defining lineage potential and fate behavior of precursors during pancreas development. Sznurkowska MK, Hannezo E, Rulands S, Nestorowa S, Hindley CJ, Azzarelli R, Nichols J, Göttgens B, Huch M, *Philpott A and *Simons BD. (2018) Developmental Cell. Aug 6;46(3):360-375. doi: 10.1016/j.devcel.2018.06.028. * Joint corresponding authors.
Subcellular localisation modulates ubiquitylation and degradation of Ascl1. Gillotin S, Davies JD, Philpott A. (2018) Sci Rep. Mar 15;8(1):4625. doi: 10.1038/s41598-018-23056-4.
Neurogenin3 phosphorylation controls pancreatic endocrine differentiation and maintenance of β-cell function. Azzarelli R, Hurley C, Sznurkowska M, Gamper I, Ali F, McCracken L, Hindley C, McDuff F, Hardwick L, Jones, Kemp R, Simons B, Huch M, Evan G, Winton D and Philpott A. (2017). Developmental Cell 41: 274-286.
Multi-site phospho-regulation of proneural transcription factors in development and reprogramming. Philpott A (2015). Neurogenesis. 2: e1049733
Ascl1 phospho-status regulates neuronal differentiation in a Xenopus developmental model of neuroblastoma. Wylie LA, Hardwick LJA, Papkovskaia TD, Thiele CJ and Philpott A (2015). Dis Model Mech. 8:429-41.
Lineage selection and plasticity in the intestinal crypt. Philpott A and Winton DJ (2014). Curr Opinion Cell Biol. Jul 29;31C:39-45.
Nervous decision-making: to divide or differentiate. Hardwick L and Philpott A (2014). Trends Genet. 30, 254-261.
The phosphorylation status of Ascl1 is a key determinant of neuronal differentiation and maturation in vivo and in vitro. Ali FR, Cheng K, Kirwan P, Metcalfe S, Livesey FJ, Barker RA and Philpott A (2014). Development. 141, 2216-24.
The cell cycle and pluripotency. Hindley C and Philpott A (2013). Biochemical Journal 451, 135-43.
Post-translational modification of Ngn2 differentially affects transcription of distinct targets to regulate the balance between progenitor maintenance and differentiation. Hindley C, Ali F, McDowell G, Cheng K, Jones A, Guillemot F and Philpott A (2012). Development,139, 1718-23.
Complex regulation controls Neurogenin3 proteolysis. Roark R, Itzhaki L and Philpott A (2012). Biology Open, 1264-72.
Regulation of cell fate determination by Skp1-Cullin1-f Box (SCF) E3 ubiquitin ligases. *Hindley CJ, McDowell GS, Wise H, Philpott A (2011). International Journal of Developmental Biology. 55:249-60.
Cell cycle-regulated multi-site phosphorylation of Neurogenin 2 coordinates cell cycling with differentiation during neurogenesis. Ali F, Hindley C, McDowell G, Deibler R, Jones A, Kirschner M, Guillemot F and Philpott A (2011). Development 138, 4267-77.

